Deep learning can be used to solve many issues in the medical field. It can make medical diagnosis' faster, more accurate, and offer better treatment solutions. With readily available X-ray data you can train a model to predict specific health conditions based on the image. In this case study, we examine a deep learning project that aims to predict health conditions in the chest from an open source of chest x-rays made publically available.
Problem
Chest X-ray exams are one of the most frequent and cost-effective medical imaging examinations available. However, clinical diagnosis of a chest X-rays can be challenging and sometimes more difficult than diagnosis via chest CT imaging. The lack of large publicly available datasets with annotations means it is still very difficult, if not impossible, to achieve clinically relevant computer-aided detection and diagnosis (CAD) in real world medical sites with chest X-rays. One major hurdle in creating large X-ray image datasets is the lack of resources for labeling so many images. Prior to the release of this dataset, Openi was the largest publicly available source of chest X-ray images with 4,143 images available.
This NIH Chest X-ray Dataset is comprised of 112,120 X-ray images with disease labels from 30,805 unique patients. To create these labels, the authors used Natural Language Processing to text-mine disease classifications from the associated radiological reports. The labels are expected to be >90% accurate and suitable for weakly-supervised learning. The original radiology reports are not publicly available but you can find more details on the labeling process in this Open Access paper: "ChestX-ray8: Hospital-scale Chest X-ray Database and Benchmarks on Weakly-Supervised Classification and Localization of Common Thorax Diseases." (Wang et al.)
To do so, we decided to tackle the ChestX-ray Kaggle challenge as a Computer Vision project, containing more than 100,000 images of size 2000x2000 pixels, which represents an overall of 50 giga image dataset.
Data
The ChestX-ray Kaggle is a challenging heavy, imbalanced and non-uniform dataset. Each 2000x2000 image is identified by a serial number and referenced in an excel file with its corresponding diagnosis estimated through radiologist report NLP analysis. Given the unbalanced distribution among the 15 different labels, the size and the variance of the chests Xray images, the preprocessing is going to be very highly computationally expensive. Hence, the use of cnvrg.io is a main requirement to tackle this project.
The cnvrg.io platform is much like a git system built for managing heavy datasets. To access the data on the platform, it needs to be pushed from a local computer. Once the data was “on the cloud” we were ready to go from our local computers to powerful machines with enough computational power to support the training of our networks.
Tasks
We used Keras and built a complete classification pipeline from loading and organizing the data to evaluating the performance:
- Binary classification that consist in building a model to discriminate between two selected classes, in our case between ‘Sick’ or ‘Healthy’ or ‘Healthy’ and some specific disease like ‘Pneumothorax’.
- Multiclass classification: training the network to be able to recognize a unique disease in the Xray among the 14 possibilities.
- Multilabel classification: be able to recognize a number of disease in the image that could occur together in the same Xray.
Challenges
One of the main issues in medical imaging analysis lies in the complexity of the image by itself. First, most of the explanatory features are obviously very small and may require several years of experience for a radiologist to detect them. In addition to this challenge, the model can be easily misled by the large discrepancy among the skeletons, the shade of colors or the image slicing. Besides the data imbalancing, the model may naively output every time the largest category present in the training set.
This unbalanced dataset is a classical problem faced when training deep learning/ machine learning models with real data. The first step would be to train beyond the naive model that will always choose the largest category present in the training set. We needed to think of different data augmentation techniques that will be discussed later.
Another main problem came with the multiclass and multilabel classification. Those are two specific problems and have led us to consider the loss functions in order to adapt the training process to each one of them.
Processing
Also, during the implementation itself we realized that we needed something more flexible than the original Keras one we wrote. Indeed, in the beginning we used the Keras library so we could use the built-in checkpoints model saver and the Image Generator to deal with the batch data flow during training. However, the need of specific data augmentation made us realize that we needed to code a more specific implementation to bypass some of the built-in tools.
To overcome the data augmentation challenge, we built our own generator function to use inside the Keras fit_generator. Each image which was underrepresented in the dataset went through ten random image augmentation functions (e.g: shift, brightness, ...). This approach has avoided deleting lot of data and has led to significantly improve our results.

Training
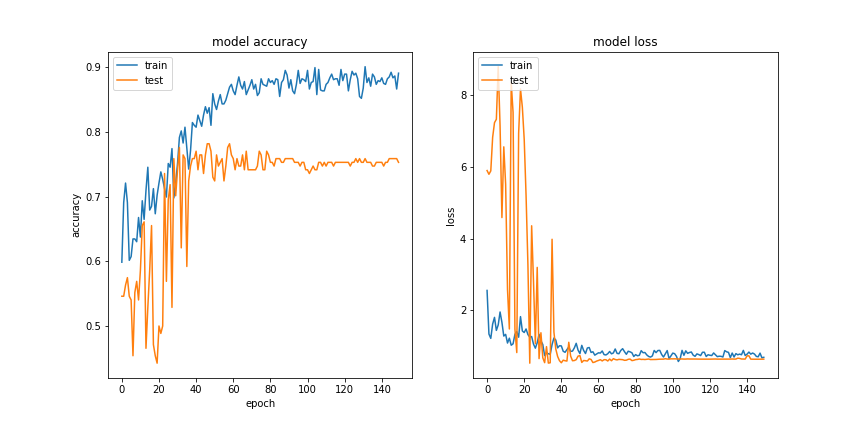
Results
These two graphs below represent the ROC curve for binary tasks. The AUC is well above 50% in both and indicates that the models are performing above random level.
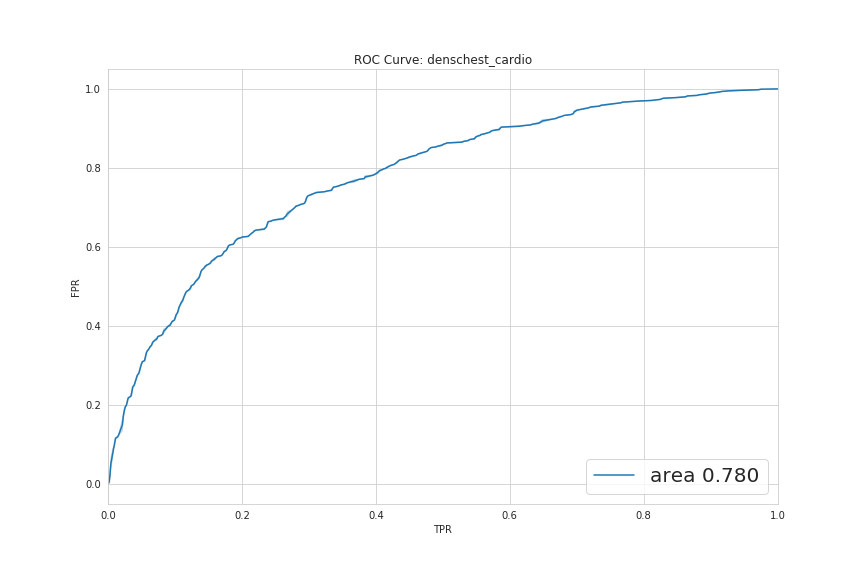
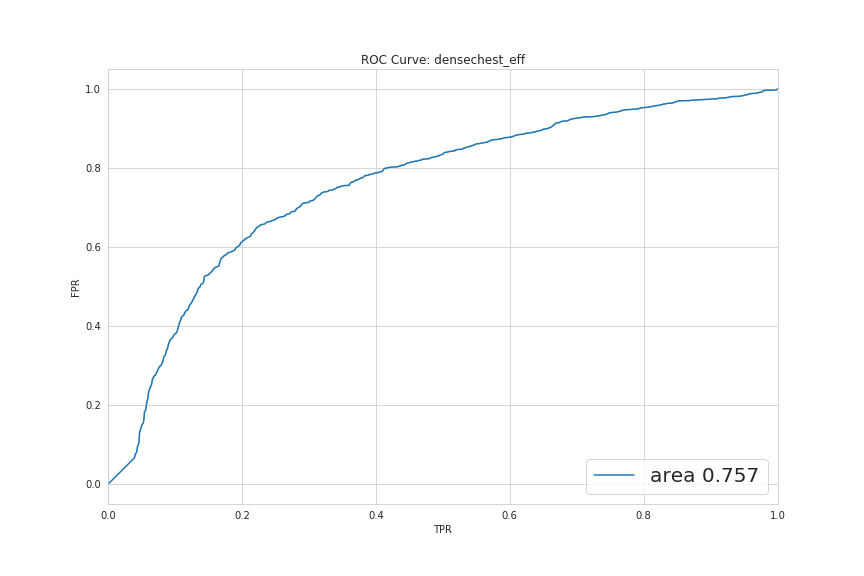
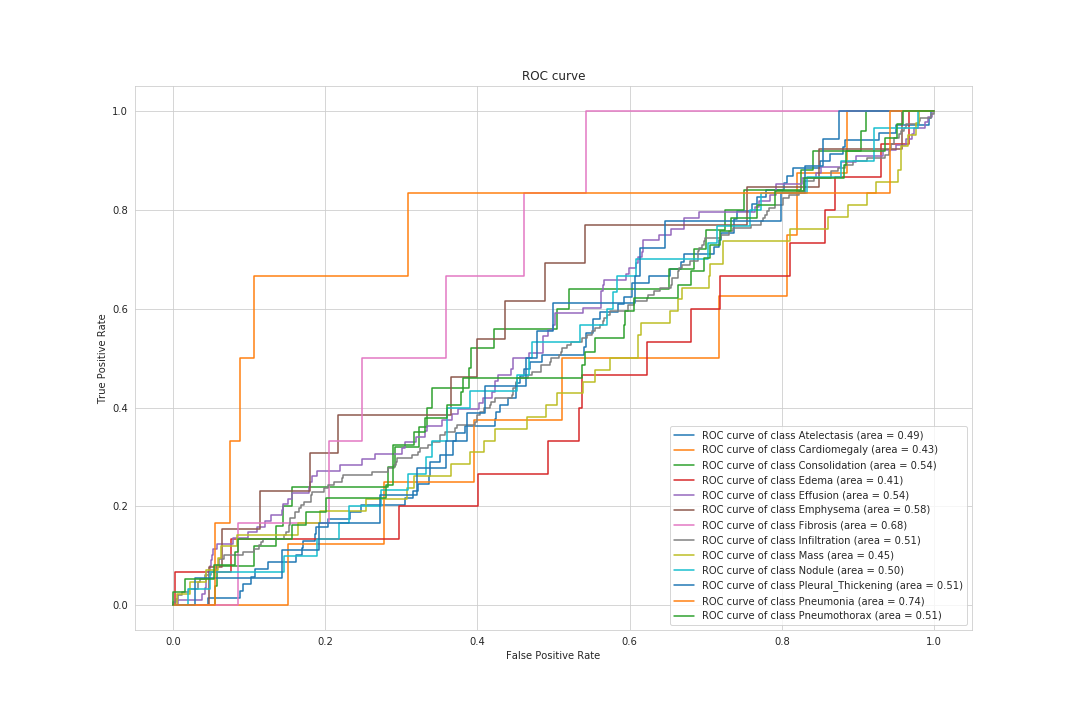
The ROC curve for the broader task of multilabel classification is showing that, for some diseases, the model performs very well (like Pneumonia) and for others, even below random.
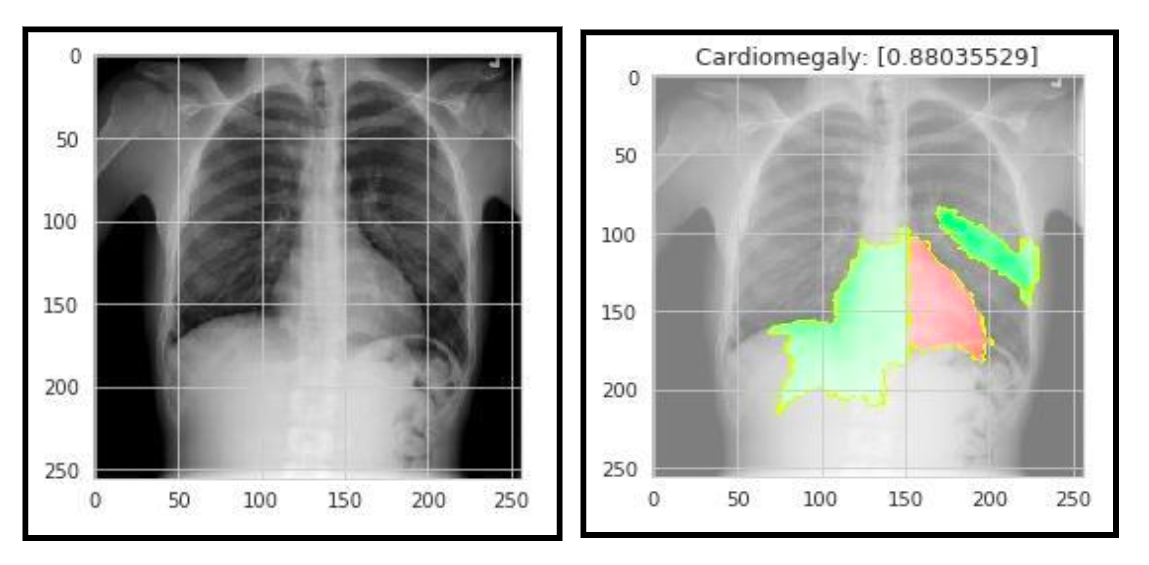
(Generated by Lime Package)
About the authors
The Chest X-ray deep learning solution was built by ITC Data Science Fellow graduates Michaël Allouche, Yair Hochner, Benjamin Lastmann, and Jeremy Eskenazi. The solution was a part of their final project for the ITC program which was taught using the cnvrg.io platform. For more information about the ITC Data Science Fellowship program, and its collaboration with cnvrg.io read this post: Building young data scientists minds, one model at a time



